The sample images I’m using in this mini tracing tutorial were of tissue sectioned into 100um-thick slices, (mounted, and tagged) in which DNA was labelled with DAPI (405nm), and we had stained for the immediate early gene (IEG) zif268 (Alexa 647), and retrovirally GFP-labelled cells (Alexa 488). DETAILS ON THE SAMPLE NEURON IMAGES USED IN THIS TUTORIAL With a 2D view, this could just seem like a shorter branch, but looking at the 3D view, if the branch ends at the edge of the z plane, it’s possible that the branch is actually longer but was just included in a different section of tissue. For example, sometimes branches that are part of the neuron are cut off the slice. When tracing neurons imaged using a confocal, it helps to keep the z plane and the overall 3D shape in mind as it could interfere with some measurements like overall dendritic length. With this, the series of 2D photos can be combined, allowing for reconstruction and 3D visualization and analyses of the specimen in question. If the 2-dimensional slices are made of the x and y planes, the thickness that is available as a third-dimension is the z plane. A series of clear 2D images can then be taken across the z plane. 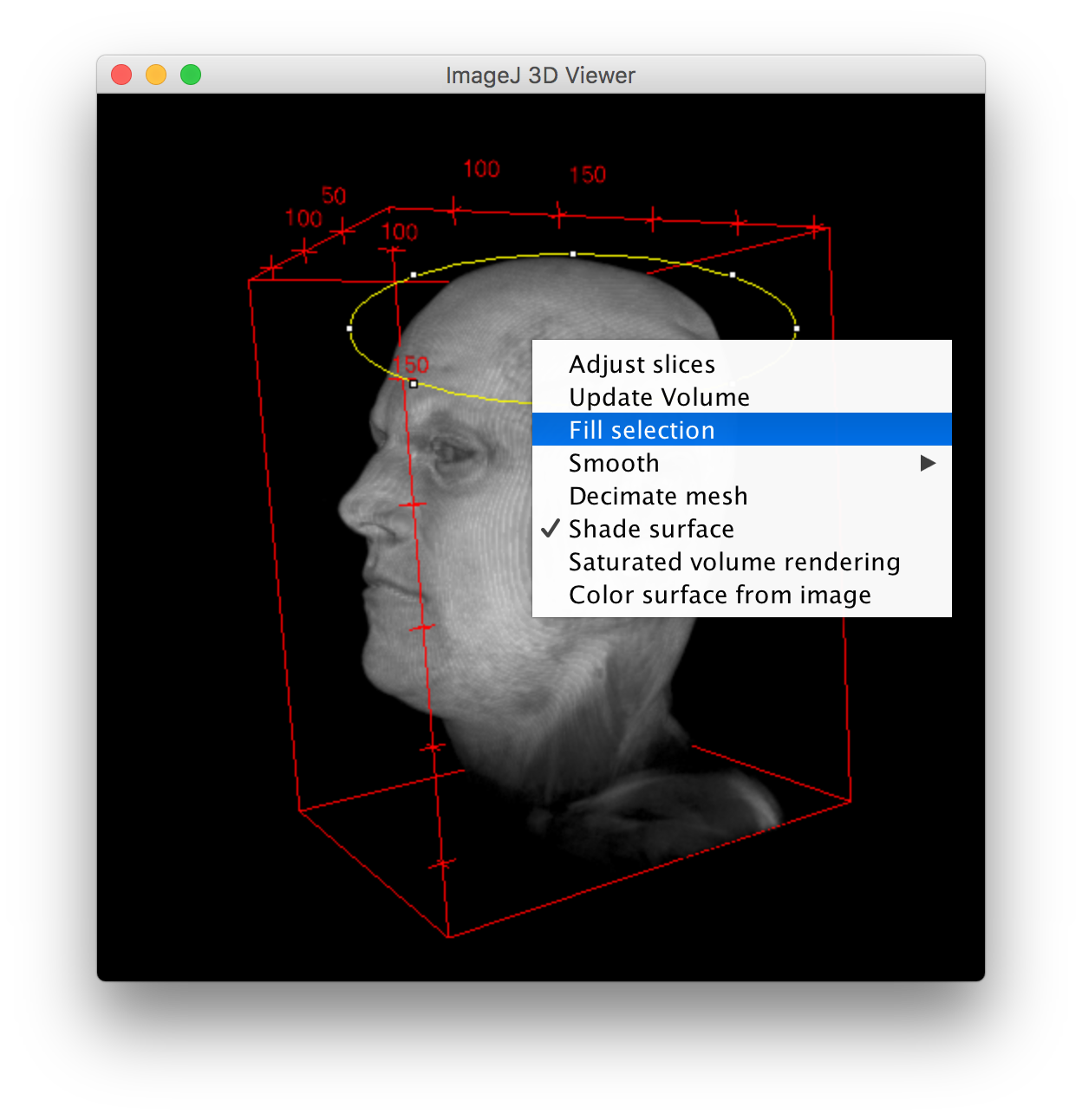
Using fluorescent imaging technique (*) and an extra spatial filter, this confocal can selectively collect light at a specific z-plane, filtering out any out-of-focus illumination. The image files used in this tutorial were taken using a Leica confocal microscope (model SP8). This post will show you how you can get one of these (dendrite tracing) using FIJI: a how-to FIJI on neuron tracing CONFOCAL IMAGING
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
